flowchart LR
A["Step 1 (Python)"] -->|Pass data as Zarr or HDF5| B["Step 2 (R)"]
Wrapping vs Native Implementation for Cross-Language Interoperability
Case study of the rhdf5 and Rarr Bioconductor packages
February 25, 2026
Intro to formats
- Competing formats to store multi-dimensional arrays (e.g., images, cell-by-gene matrices, etc.) on disk
- Particularly important for large datasets that don’t fit in memory
- Basis of interoperability between languages (e.g., R, Python, Julia, etc.)
Zarr vs HDF5: technical
Zarr is “cloud-native”, and easily parallelizable.
For both Zarr and HDF5, chunks of the file can be accessed.
BUT if the file is stored remote, it needs to be downloaded first for HDF5, while Zarr can be fetch only the relevant chunk.

/tmp/Rtmp1aeF1J/file3f939f999db.zarr
├── 0
│ ├── 0
│ ├── 1
│ └── 2
├── 1
│ ├── 0
│ ├── 1
│ └── 2
└── 2
├── 0
├── 1
└── 2Zarr vs HDF5: governance
Zarr is community-driven:
- much younger
- less normative(?) (at least for now): e.g., sparse arrays
- relies more on “subformats”(?): OME
Definitions
- Wrapping: using a high-level language (e.g., R) to call functions from an underlying library (e.g., C/C++/Python) that implements the spec.
- Vendoring: including a copy of the underlying library in the package source code, and wrapping it in the high-level language of choice.
- Native implementation: implementing the spec from scratch, without including any third-party code in the package source code.
rhdf5 overview
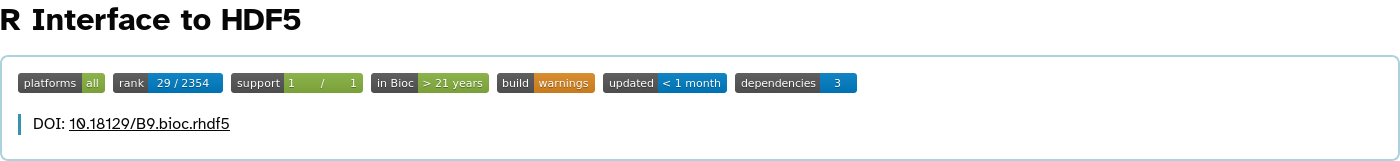

Rarr overview
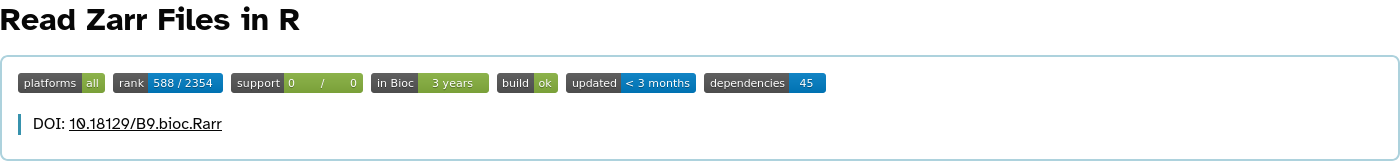

Vendoring: rhdf5 case
Wrapping and vendoring the HDF5 C library.
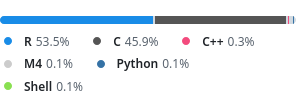
Lots of C code, but also lots of thin wrappers handling R memory management and R/C data type conversions.
Native implementation: Rarr case
Native implementation of the Zarr spec in R.
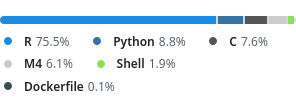
Most prep steps and housekeeping is done in R. Only performance critical steps are in C.
These steps should eventually also run in parallel or on GPU.
Aside / caveat
Zarr specification is still “python-biased” and includes many “numpysms”.
Better since version 3
- Some compression libraries are bundled in Rarr
Wrapping vs Native implementation
Shared problems:
- Upgrading to a new version is always very costly
- Everybody keeps releasing competing / overlapping packages
Wrapping vs Native implementation
Pros wrapping:
Faster to get an initial proof of concept.
No need to start from scratch.
- Potentially better maintained & optimized underlying library
- Lower risk of spec misinterpretation or misimplementation
Mitigations: conformance testing
Mitigations: conformance testing
Zarr conformance tests:
- compare consistency across implementations
- benchmark performance across implementations
- identify edge cases and spec ambiguities
- provide a high number of test cases “for free” to new implementations
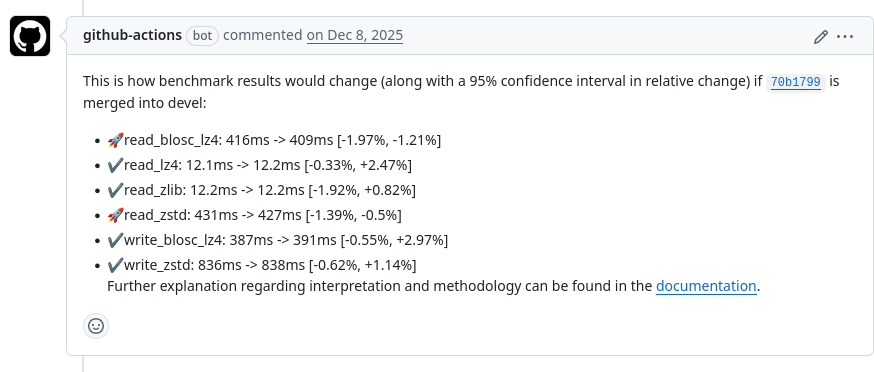
Mitigations: continuous benchmarking

Request for help
Low diversity of test datasets.
Artür discovered a lot of edge cases and missing features when implementing Zarr support in anndataR.
Vendoring vs Native implementation
Cons vendoring (1/3):
- Clash with CRAN / Bioconductor policies
* checking compiled code ... WARNING Note: information on .o files is not available File ‘/home/biocbuild/bbs-3.22-bioc/R/site-library/rhdf5/libs/rhdf5.so’: Found ‘__sprintf_chk’, possibly from ‘sprintf’ (C) Found ‘abort’, possibly from ‘abort’ (C) Found ‘rand_r’, possibly from ‘rand_r’ (C) Found ‘stderr’, possibly from ‘stderr’ (C) Found ‘stdout’, possibly from ‘stdout’ (C) Compiled code should not call entry points which might terminate R nor write to stdout/stderr instead of to the console, nor use Fortran I/O nor system RNGs nor [v]sprintf. The detected symbols are linked into the code but might come from libraries and not actually be called.
Vendoring vs Native implementation
Cons vendoring (2/3):
- Larger library (size & API) than needed
- is manually sorting through the files worth it?
- Patching process:
- git patch vs script
- applied to vendored version vs applied at build time
- Underlying library influence wrapper API = less idiomatic API
Vendoring vs Native implementation
Cons vendoring (3/3):
- Clash with Debian policy
- Lower control on update timing
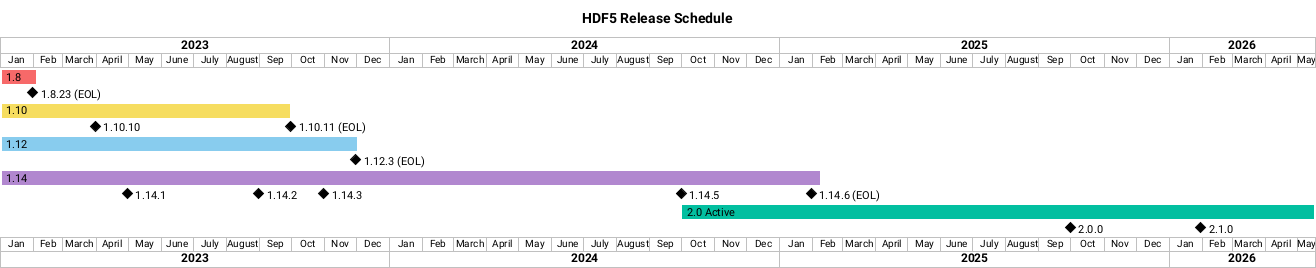

Support for older versions: challenges
By vendoring, we can make sure all users have the same version.
vs wrapping a system library:

Support for older versions: policy
- We always support reading from older formats
- Support for writing to older formats is done on a best effort basis
Making updates easier
Convince libraries to stick more closely to semantic versioning
Now the case in HDF5 2.0.0!
- Pay technical debt early: “Frequency Reduces Difficulty”
Conclusion

Hugo Gruson; Huber Group Lab Meeting 02/2026